Charvet et al., medRxiv, 2022.01
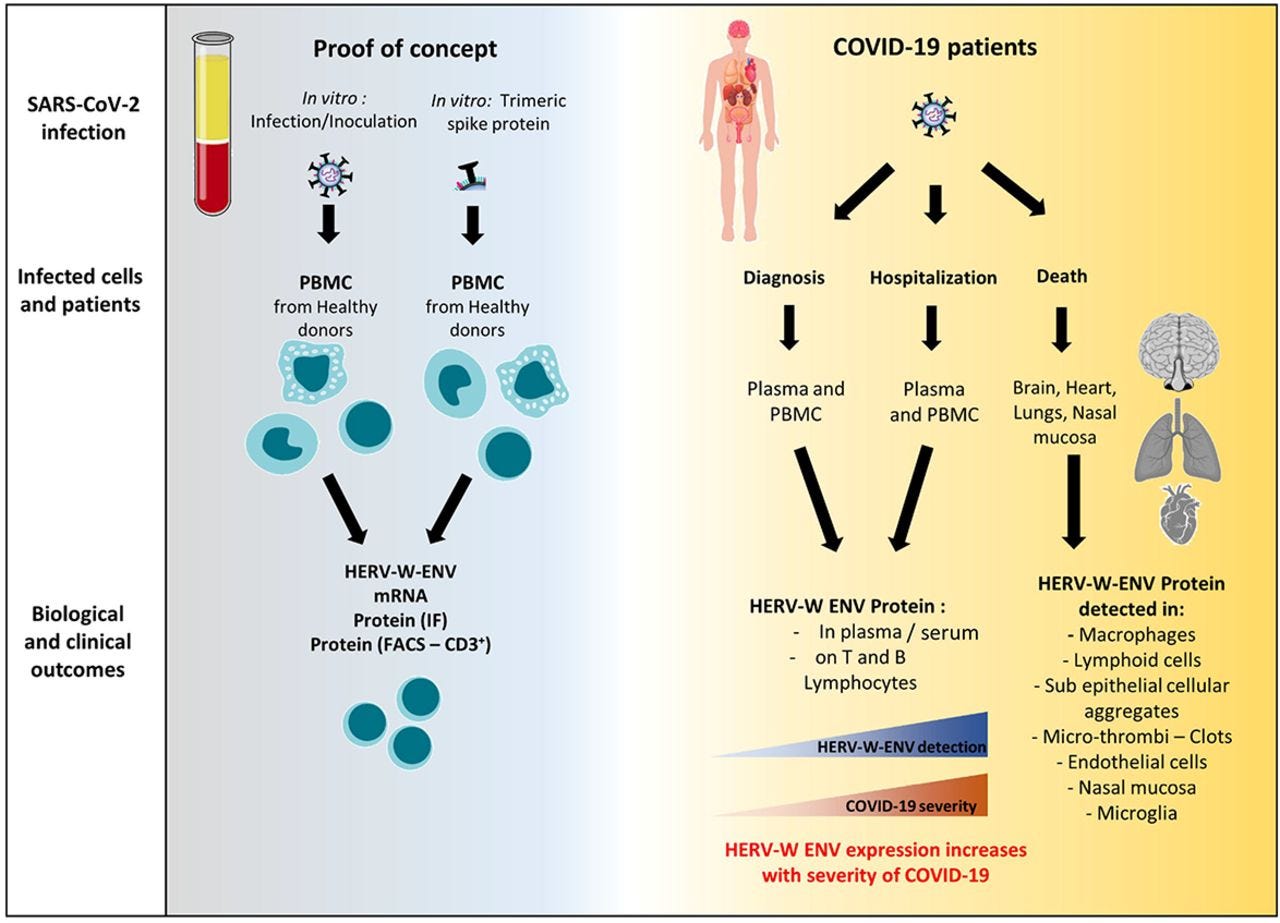
“Patients with COVID-19 may develop abnormal inflammatory response and lymphopenia, followed in some cases by delayed-onset syndromes, often long-lasting after the initial SARS-CoV-2 infection. As viral infections may activate human endogenous retroviral elements (HERV), we studied the effect of SARS-CoV-2 on HERV-W and HERV-K envelope (ENV) expression, known to be involved in immunological and neurological pathogenesis of human diseases. Our results have showed that the exposure to SARS-CoV-2 virus activates early HERV-W and K transcription but only HERV-W ENV protein expression, in an infection- and ACE2-independent way within peripheral blood mononuclear cell cultures from one-third of healthy donors. Moreover, HERV-W ENV protein was significantly increased in serum and plasma of COVID-19 patients, correlating with its expression in CD3+ lymphocytes and with disease severity. Finally, HERV-W ENV was found expressed in post-mortem tissues of lungs, heart, brain olfactory bulb and nasal mucosa from acute COVID-19 patients in cell-types relevant for COVID-19-associated pathogenesis within affected organs, but different from those expressing of SARS-CoV-2 antigens. Altogether, the present study revealed that SARS-CoV-2 can induce HERV-W ENV expression in cells from individuals with symptomatic and severe COVID-19. Our data suggest that HERV-W ENV is likely to be involved in pathogenic features underlying symptoms of acute and post-acute COVID. It highlights the importance to further understand patients’ genetic susceptibility to HERV-W activation and the relevance of this pathogenic element as a prognostic marker and a therapeutic target in COVID-19 associated syndromes.”
SARS-CoV-2 Proteins Bind to Hemoglobin and Its Metabolites
Lechuga et. al, International Journal of Molecular Sciences, 2021.08
“Severe COVID-19, caused by SARS-CoV-2, typically leads to pneumonia and acute respiratory distress syndrome (ARDS). However, the growing list of evidence indicates a systemic impairment that leads to multiorgan failure. During an infection, an imbalance in the immunological response can produce a “cytokine storm” and numerous other pathophysiological processes such as hypoxemia, thrombosis, pulmonary embolism, encephalopathy, myocardial injury, heart failure, and acute kidney injury. Hematological dysfunction in severe COVID-19 includes low levels of erythrocytes and an increased variation in the red blood cell distribution width (RDW). Recent reports on the immune effects of COVID-19 have highlighted immune thrombocytopenia and autoimmune hemolytic anemia. Other evidence suggests an increase in hemophagocytosis related to elevated ferritin levels in COVID-19. In hemolytic disorders, the release of high levels of Hb and heme triggers the pro-inflammatory response, complement activation, and procoagulant and pro-oxidative environment.”
“Overall, our demonstration that SARS-CoV-2 proteins can bind to heme or Hb may have clinical implications. Mainly, Hb’s interaction with spike opens new therapeutic perspectives due to significant virus attachment and replication inhibition. Moreover, this binding could potentially increase or drive hematological disorders and thrombosis observed in severe COVID-19. Although there are still knowledge gaps on viral-host cell complex interplay and disease pathophysiology, the data presented here will contribute to scientific discussion.”
“Conclusions: In this study, we identified that (at least) five proteins (S, N, M, Nsp3, and Nsp7) of SARS-CoV-2 recruit Hb/metabolites. The motifs of the RDB of SARS-CoV-2 spike, which binds Hb, and the sites of the heme bind-N protein were disclosed. In addition, these compounds and PpIX block the virus’s adsorption and replication. Furthermore, we also identified heme-binding motifs and interaction with hemin in N protein and other structural (S and M) and non-structural (Nsp3 and Nsp7) proteins.”
Fig. 3. “Hemoglobin (Hb) binding to spike protein and RBD enhancement. (a) Total protein extracts (20 μg) of SARS-CoV- 2-infected Vero E6 cells (Infected) or mock-infected cells (Mock; 20 μg) were separated by SDS-PAGE (10%) along with haptoglobin (Hp; 2 μg), transferred to a nitrocellulose membrane, and serially incubated with Hb (10 μg/mL), anti-Hb antibody, and peroxidase-conjugated secondary antibodies. Two bands at 72 and 150 kDa were exclusively found in SARS-CoV-2 infected extract (arrowhead). (b) Spot synthesis analysis with a library of 15-mer peptides offset by five amino acids to represent the RBD region (purple) and NTD (blue) of the spike protein, haptoglobin (Hapt; red), and casein (green) as a positive and negative control, respectively. Sequences were synthesized directly onto a cellulose membrane followed by probing with Hb (5 μg/mL) and revealed by anti-human Hb antibodies. The top panel shows the chemiluminescent image of signals from peptides bound to Hb. The bottom panel shows a graph of the signal intensities normalized to the maximum signal. An intensity level above 50% defined Hb-reactive peptides. Molecular docking of SARS-CoV-2 spike protein with human hemoglobin. (c) Interaction of spike protein (gray) with α-chain Hb (orange) and β-chain (red). (d) Representation of amino acid residues binding to spike protein (sticks and gray) with α-chain of hemoglobin (orange).”
Zhang et al., PNAS, 2021.05
“An unresolved issue of SARS-CoV-2 disease is that patients often remain positive for viral RNA as detected by PCR many weeks after the initial infection in the absence of evidence for viral replication. We show here that SARS-CoV-2 RNA can be reverse-transcribed and integrated into the genome of the infected cell and be expressed as chimeric transcripts fusing viral with cellular sequences. Importantly, such chimeric transcripts are detected in patient-derived tissues. Our data suggest that, in some patient tissues, the majority of all viral transcripts are derived from integrated sequences. Our data provide an insight into the consequence of SARS-CoV-2 infections that may help to explain why patients can continue to produce viral RNA after recovery.”
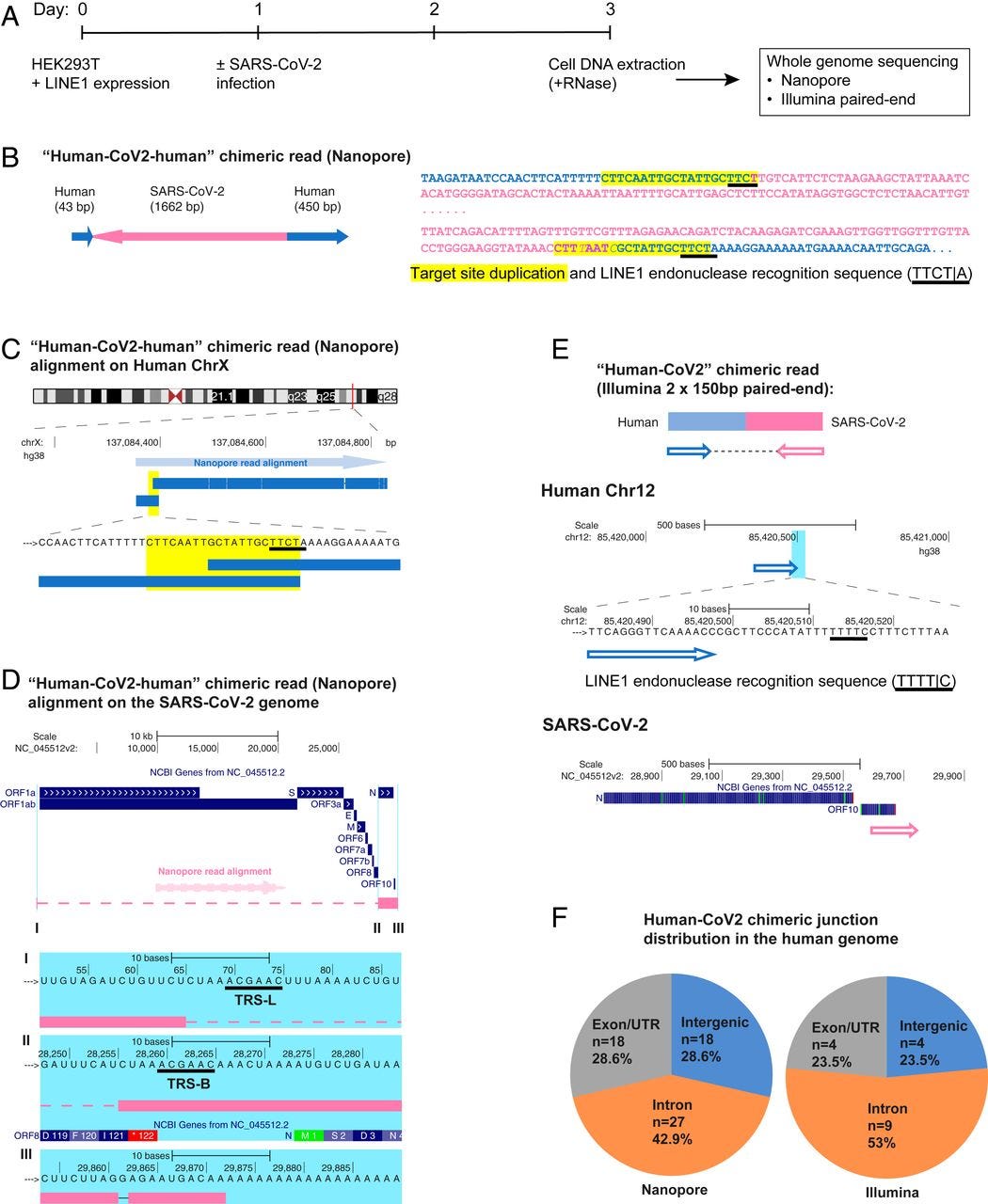
Fig. 1 “SARS-CoV-2 RNA can be reverse transcribed and integrated into the host cell genome. (A) Experimental workflow. (B) Chimeric sequence from a Nanopore sequencing read showing integration of a full-length SARS-CoV-2 NC subgenomic RNA sequence (magenta) and human genomic sequences (blue) flanking both sides of the integrated viral sequence. Features indicative of LINE1-mediated “target-primed reverse transcription” include the target site duplication (yellow highlight) and the LINE1 endonuclease recognition sequence (underlined). Sequences that could be mapped to both genomes are shown in purple with mismatches to the human genomic sequences in italics. The arrows indicate sequence orientation with regard to the human and SARS-CoV-2 genomes as shown in C and D. (C) Alignment of the Nanopore read in B with the human genome (chromosome X) showing the integration site. The human sequences at the junction region show the target site, which was duplicated when the SARS-CoV-2 cDNA was integrated (yellow highlight) and the LINE1 endonuclease recognition sequence (underlined). (D) Alignment of the Nanopore read in B with the SARS-CoV-2 genome showing the integrated viral DNA is a copy of the full-length NC subgenomic RNA. The light blue highlighted regions are enlarged to show TRS-L (I) and TRS-B (II) sequences (underlined, these are the sequences where the viral polymerase jumps to generate the subgenomic RNA) and the end of the viral sequence at the poly(A) tail (III). These viral sequence features (I–III) show that a DNA copy of the full-length NC subgenomic RNA was retro-integrated. (E) A human–viral chimeric read pair from Illumina paired-end whole-genome sequencing. The read pair is shown with alignment to the human (blue) and SARS-CoV-2 (magenta) genomes. The arrows indicate the read orientations relative to the human and SARS-CoV-2 genomes. The highlighted (light blue) region of the human read mapping is enlarged to show the LINE1 recognition sequence (underlined). (F) Distributions of human–CoV2 chimeric junctions from Nanopore (Left) and Illumina (Right) sequencing with regard to features of the human genome.”
The Global Phosphorylation Landscape of SARS-CoV-2 Infection
Bouhaddou et al., Cell, 2020.08
Highlights
Phosphoproteomics analysis of SARS-CoV-2-infected cells uncovers signaling rewiring
Infection promotes host p38 MAPK cascade activity and shutdown of mitotic kinases
Infection stimulates CK2-containing filopodial protrusions with budding virus
Kinase activity analysis identifies potent antiviral drugs and compounds
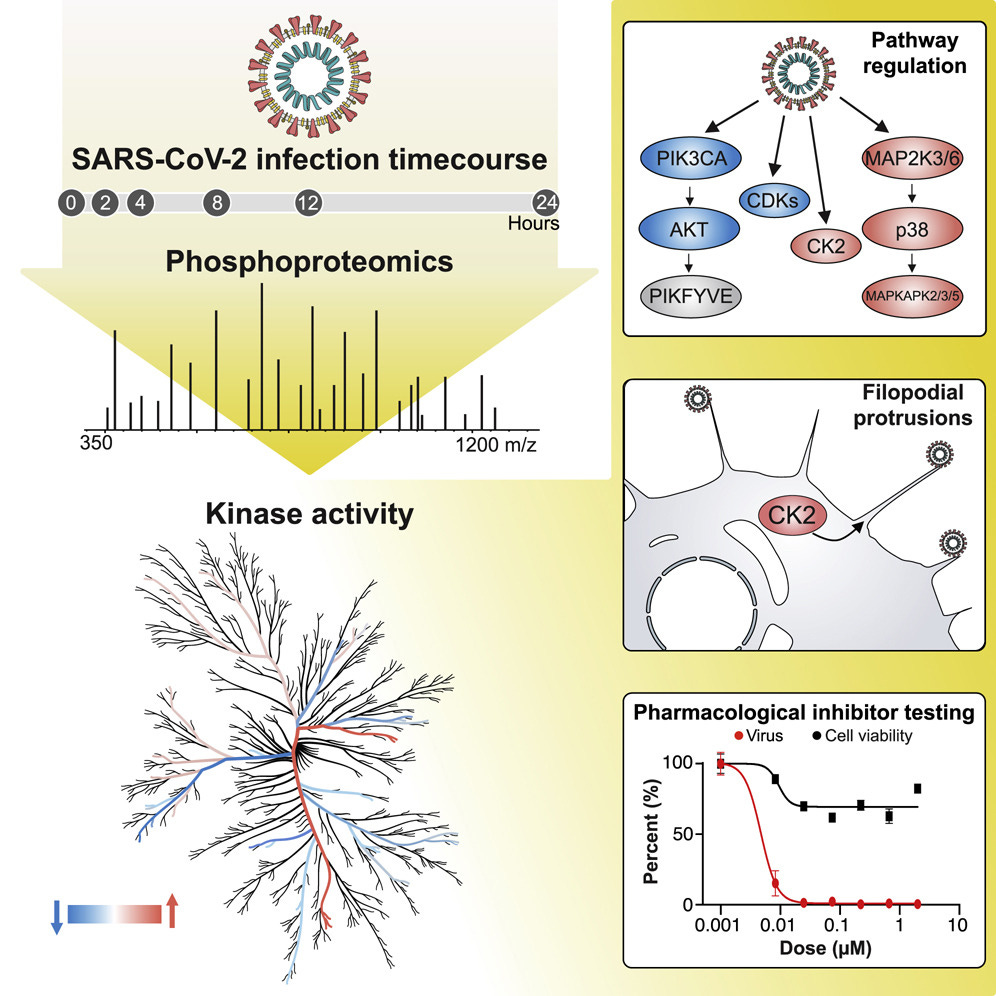
“The causative agent of the coronavirus disease 2019 (COVID-19) pandemic, severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), has infected millions and killed hundreds of thousands of people worldwide, highlighting an urgent need to develop antiviral therapies. Here we present a quantitative mass spectrometry-based phosphoproteomics survey of SARS-CoV-2 infection in Vero E6 cells, revealing dramatic rewiring of phosphorylation on host and viral proteins. SARS-CoV-2 infection promoted casein kinase II (CK2) and p38 MAPK activation, production of diverse cytokines, and shutdown of mitotic kinases, resulting in cell cycle arrest. Infection also stimulated a marked induction of CK2-containing filopodial protrusions possessing budding viral particles. Eighty-seven drugs and compounds were identified by mapping global phosphorylation profiles to dysregulated kinases and pathways. We found pharmacologic inhibition of the p38, CK2, CDK, AXL, and PIKFYVE kinases to possess antiviral efficacy, representing potential COVID-19 therapies.”
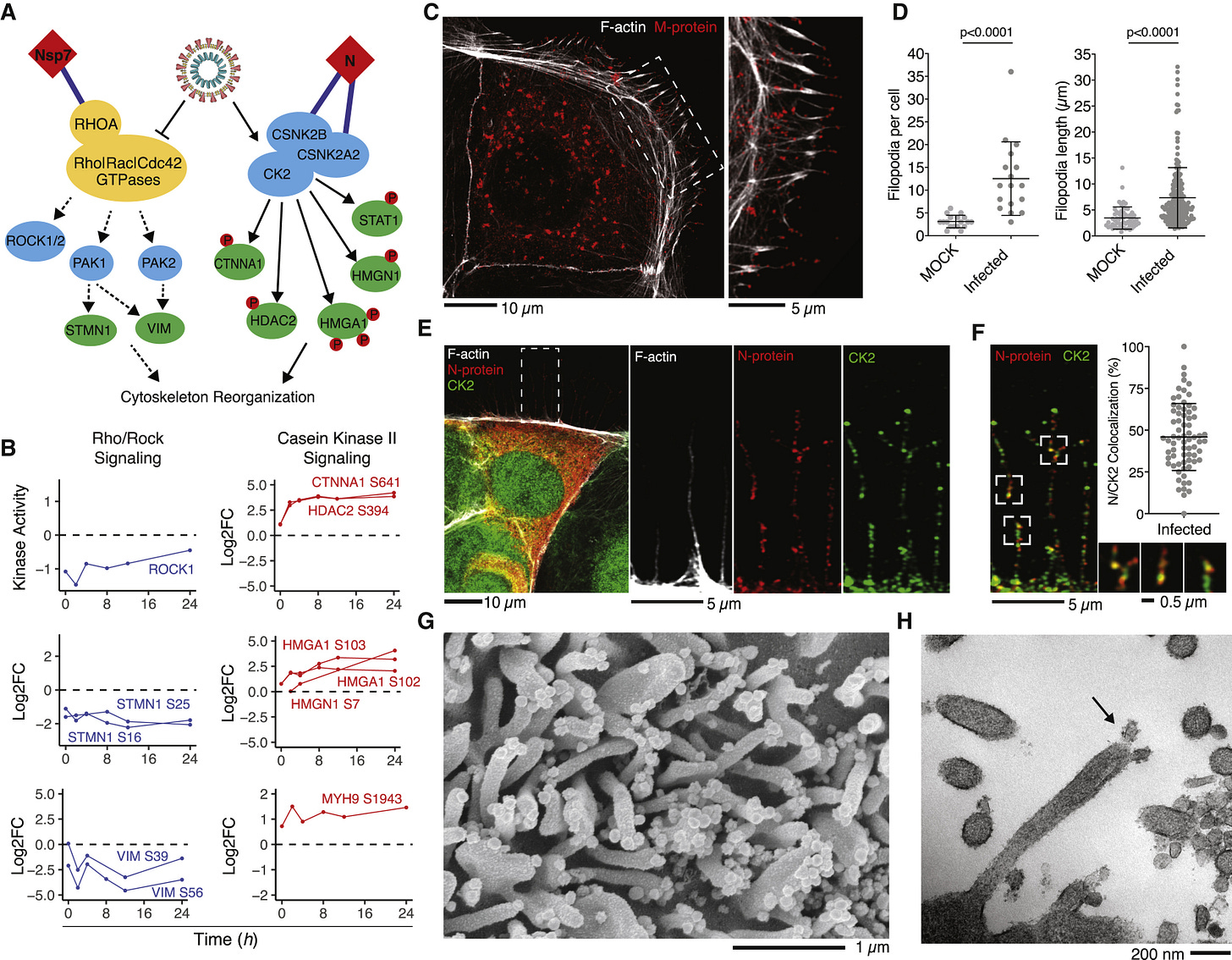
Fig. 5. Colocalization of CK2 and Viral Proteins at Actin Protrusions
(A) Pathway of regulated PHs and SARS-CoV-2 interaction partners involved in cytoskeletal reorganization. Dashed lines indicate downregulation of activity, while solid lines indicate upregulation of activity. (B) Regulation of individual kinase activity or PHs depicted in (A). (C) Caco-2 cells infected with SARS-CoV-2 at an MOI of 0.1 for 24 h prior to immunostaining for F-actin and M protein, as indicated. Shown is a confocal section revealing M protein localization along and to the tip of filopodia (left) and magnification of the dashed box (right). (D) Dot plot quantification of the number and length of filopodia in untreated (mock) or infected Caco-2 cells for 24 h with SARS-CoV-2. Filopodium length was measured from the cortical actin to the tip of the filopodium. Error bars represent SD. Statistical testing by Mann-Whitney test. (E) Caco-2 cells infected with SARS-CoV-2 at an MOI of 0.01 for 24 h prior to immunostaining for F-actin, N protein, and casein kinase II (CK2) as indicated (left). Shown is magnification of the dashed box as single channels (right). (F) Magnification of the dashed box from (E) with quantification of colocalization between CK2 and N protein throughout infected Caco-2 cells. Displayed is the proportion of N protein-positive particles colocalizing with CK2. Error bars represent SD. (G and H) Scanning electron microscopy (G) and transmission electron microscopy (H) images of SARS-CoV-2 budding from Vero E6 cell filopodia (black arrow in H).